VIB research team uses AI to explain cell diversity in the fly brain
Landmark fly brain study deciphers the genomic regulatory code behind neuronal diversity
The single-cell revolution is revealing more and more fine-grained insights into the biology of living organisms. The deep learning (AI) revolution adds unprecedented computational power to analyze these “big data”. A research team led by Stein Aerts (VIB-KU Leuven) has combined these two revolutions to sketch a new picture of gene regulation in all cells of the fruit fly brain, using deep learning to reveal how specific pieces of DNA steer neuronal identity, from birth to maturity. This landmark achievement, published today in Nature, is one of the signposts of a new era of AI-driven biomedical research with unprecedented possibilities, including personalized cell-based interceptive medicine.
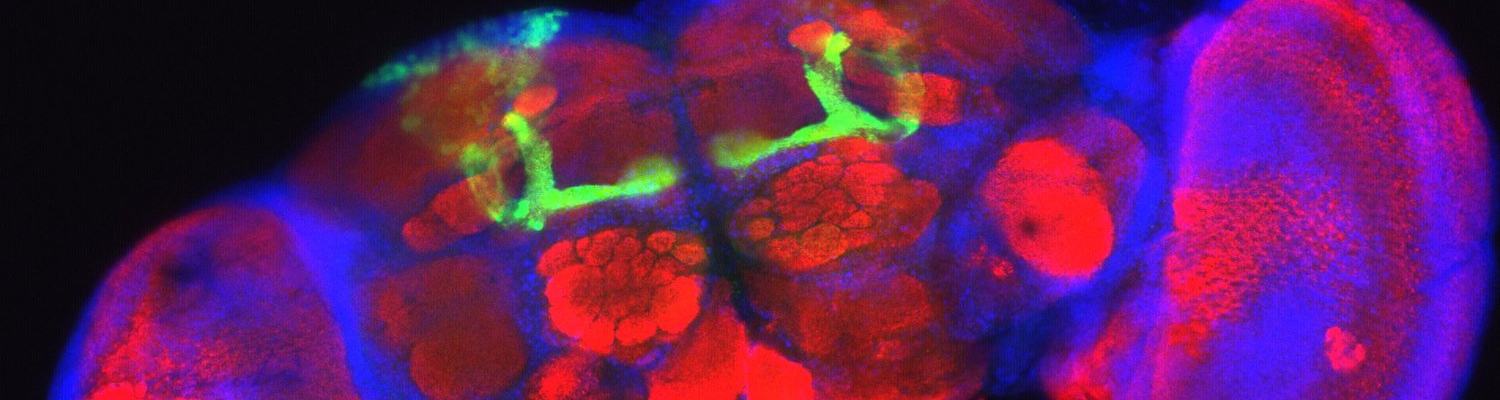
All the cells in our body share the same genetic code or DNA. Still, these cells have widely different shapes and functions, from muscle cells to fat cells or brain cells. “Particularly within the brain, the most complex organ, there is an immense diversity of neuronal and glial cell types that are wired together in a circuit to enable a broad range of functional and behavioral traits,” explains prof. Stein Aerts of the VIB-KU Leuven Center for Brain & Disease Research. “This is true for us humans, but also for tiny animals such as the fruit fly.”
Aerts’ team is specialized in decoding how our DNA is used to create all the different cell types and how it drives dynamic changes in cellular states—not only in human tissue but also in fruit flies. Aerts: “The fruit fly brain contains ‘only’ around 220,000 cells, a scale for which it has now become feasible to investigate the full diversity of all cell types.”
Decoding DNA access
The sequence of our DNA holds all genetic information, but this is only part of the story. The long DNA helix encodes all the genes that are necessary to create different cell types, and more importantly, all the information to control which cells use which genes and when. This information is stored in short stretches of DNA, called enhancers. However, unlike the genetic code of genes, that was deciphered in the late 1960’s, the code of enhancers is still largely unknown.
“Of central importance to cell identity is the match between transcription factors and enhancers in the DNA,” explains Aerts. “Transcription factors are proteins that decode the regulatory information in our cells by recognizing enhancers, thereby making certain pieces of DNA accessible. We wanted to understand how they do that. We used a recently developed technique called single-cell assay for transposase accessible chromatin by sequencing (scATAC-seq) to identify all the enhancers that are accessible in a given cell type, at any given time.”
From larva, to pupa and adult
In a big team effort, the Aerts lab used scATAC-seq to profile DNA accessibility of 250,000 cells during brain development at larval, pupal and adult stages. The experiments were spearheaded by Jasper Janssens, Sara Aibar and Ibrahim Ihsan Taskiran, three early-career researchers in Aerts’ team.
“Our atlas showed that all cell types in the fly brain have unique DNA accessibility profiles, with tens of thousands of accessible regions per cell type,” explains Jasper Janssens. “We identified over 96,000 potential enhancers, covering about a third of the entire fly genome!”
“We then used deep learning to integrate this information on DNA accessibility with gene expression to construct what we call ‘enhancer-driven gene regulatory networks’,” adds Aibar. “These networks explain which transcription factors bind to which enhancers to regulate particular target genes. In this way, we could predict the effect of a factor on the DNA and on cell function, and trace neuronal and glial cell types from birth to maturity.”
Taskiran: “With our deep learning approach, we could reveal motifs that are missed by conventional algorithms. As such, we identified new rules on how enhancers are built and which sequences are important for their function.”
From flies to humans
“Of course, our results offer a starting point for much more fine-grained spatiotemporal genetic analysis in fruit flies,” says Aerts. “About two-thirds of the transcription factors in the networks we identified are linked to known brain defects or human disease, providing a foundation for follow-up studies. Indeed, brain development is surprisingly conserved through evolution!”
The building rules for enhancers revealed in this study can also be used to design enhancers that target a cell type at a specific timepoint, making it easier to manipulate and characterize certain cells. These novel tools will lead to a better understanding of the different neurons in the brain. To maximize the value of these new insights for the scientific community, the researchers made all data publicly available online.
According to Aerts, this fruit fly study provides an important stepping stone for a better understanding of gene regulation in humans as well: “Given the ongoing work in mouse and human tissue and further technological improvements, it is only a matter of time before similar networks will be derived for human brain cells as well. These could then be used as the starting point to design enhancers for gene therapy to treat neurodegenerative diseases for example.”
Aerts is member of a European-wide consortium aiming to track, understand and target human cells during the onset and progression of disease, and to analyze their response to therapy at single-cell resolution. “I am convinced that current advances in single-cell technology, deep learning and bioinformatics will transform how we tackle medical challenges such as cancer and brain disease in our lifetime.”
Publication
Janssens, Aibar, Ihsan Taskiran et al. Nature 2022
Decoding gene regulation in the fly brain
Questions from patients
A breakthrough in research is not the same as a breakthrough in medicine. The realizations of VIB researchers can form the basis of new therapies, but the development path still takes years. This can raise a lot of questions. That is why we ask you to please refer questions in your report or article to the email address that VIB makes available for this purpose: patienteninfo@vib.be. Everyone can submit questions concerning this and other medically-oriented research directly to VIB via this address.