Designing better membrane proteins by embracing imperfection
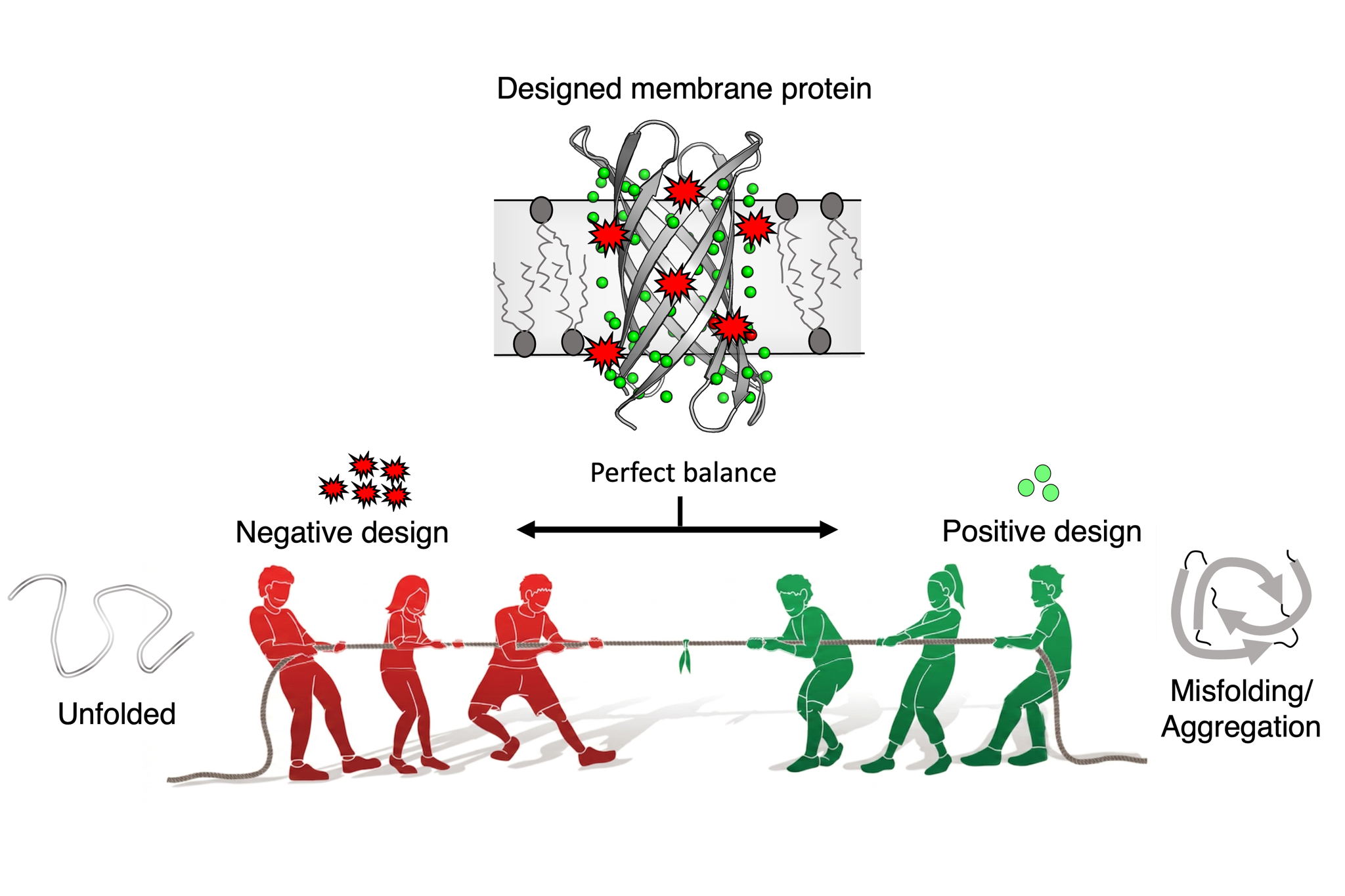
Brussels, 14 April 2026 — Scientists at the VIB–VUB Center for Structural Biology have uncovered a counterintuitive principle that could reshape how membrane proteins are designed from scratch: sometimes, making a protein less stable helps it fold correctly. In a study published in Proceedings of the National Academy of Sciences (PNAS), researchers demonstrate that introducing carefully placed ‘imperfections’, a strategy known as negative design, enables synthetic membrane proteins to fold and assemble efficiently in artificial membranes.
Membrane protein stability
Membrane proteins are essential for life and biotechnology, acting as gateways, sensors, and drug targets. Yet designing them from scratch remains notoriously difficult. Unlike soluble proteins, they must navigate a complex folding process while inserting into lipid membranes and during this step, many designs fail.
Traditional protein design focuses on maximizing the stability of the final folded structure. But the new study shows that, for transmembrane β-barrel proteins, this approach can backfire.
Using a cell-free protein synthesis system combined with synthetic lipid vesicles, the team found that highly optimized designs often misfold and aggregate instead of inserting into membranes.
“Designing for maximum stability alone can actually trap these proteins in the wrong state,” says first author and PhD student Giacomo Pedrelli (VIB-VUB). “They become too eager to fold too early, which leads to aggregation in water before they ever reach the membrane.”

The power of “negative design”
To overcome this, the researchers introduced subtle destabilizing features to disrupt premature folding. This negative design strategy reduced aggregation and significantly improved membrane insertion and assembly. Remarkably, these changes did not substantially compromise the final stability of the proteins. Instead, they helped guide the folding pathway, ensuring the protein reached the membrane in a foldable state.
The study also revealed that a protein language model (ESM3), trained on evolutionary data, outperformed traditional physics-based methods in identifying beneficial negative design mutations. While conventional tools predicted these mutations would destabilize the protein, the AI model successfully pinpointed changes that improved assembly in membranes.
The ability to reliably design transmembrane β-barrels opens exciting possibilities. These proteins can form nanopores — tiny channels with applications in biosensing, molecular detection, and next-generation sequencing technologies.
“This work shows that we need to think beyond static structures,” says Prof. Anastassia Vorobieva (VIB-VUB). “By designing not just the final state, but taking in consideration the entire folding journey, we can unlock new possibilities for engineering functional membrane proteins.”
This negative design approach for designing membrane proteins could accelerate the development of synthetic proteins for biotechnology, medicine, and nanotechnology.
Publication
Negative design enables cell-free expression and folding of designed transmembrane β-barrels. Pedrelli, et al., PNAS (2026). DOI: 10.1073/pnas.2528772123
Funding
This work was supported by Research Foundation Flanders – FWO and ERC.
Gunnar De Winter
.png)